 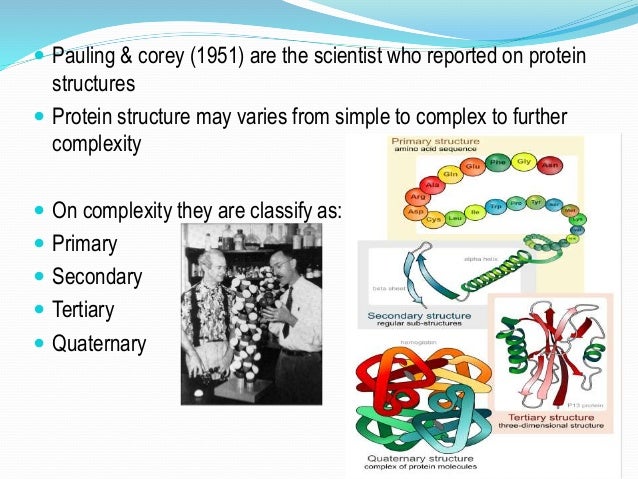
P-SEA assigns secondary structure using Cα coordinates. STRIDE is optimised to meet the visual assignments made by crystallographers. Torsion angles are given propensities according to how close they are to their regions in the Ramachandran plot. The STRuctural IDEntification method (STRIDE) uses empirically derived hydrogen bond energy and phi-psi torrsion angles to assign secondary structure.  
The continuous output is achieved by running DSSP multiple times with different hydrogen bond thresholds and then imposing a weighted average over the individual DSSP assignments to each residue. DSSPcontĭSSPcont is a variant of DSSP aiming to assign secondary structure in a continuous fashion, thus reflecting the structural variability due to thermal motion. DSSP defines a hydrogen bond where the bond energy is below -0.5 kcal/mol based on a Coulomb approximation of the hydrogen bond energy. DSSPĭictionary of Secondary Structure of Proteins (DSSP) assigns eight state secondary structure using hydrogen bonds alone. Janes, 2010, 2Struc - The Protein Secondary Structure Analysis Server, Biophysical Journal, 98:454a-455) and each of the methods you run. If you use 2Struc and publish your work please cite our paper (Klose, D & R.W. In each reference box you will find a link to the original published work and, where available, web based resources. Please click the purple buttons to access the references. Presented below is a short description of each method applied in 2Struc. 
0 Comments
Leave a Reply. |
 RSS Feed
RSS Feed
